Lab 11: PROCOTOL
Assay overview and technical considerations
Overall Experimental Setup:
You will be given two 96-well plates to complete four steps:
- A pilot experiment for DNSA binding
- A DNSA titration to determine KD for DNSA
- A pilot experiment for AZ binding
- An AZ titration to determine KD for AZ
Each plate will look something like this:
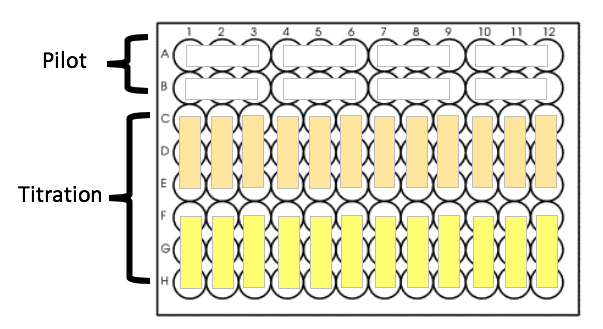
GENERAL RECOMMENDATIONS:
- Do everything in triplicate (even the pilots!), unless you do not have enough protein.
- Be sure to include a no-protein blank on every plate, and remember to collect data for both wt and mutant HCAII.
- Plan for a final protein concentration of 0.25 μM protein.
- Use multichannel pipettes whenever possible! They save significant time and reduce error.
- To make planning and pipetting easier, your final volumes can differ by up to ~5 μL.
- What this means is that you will pipette 100 μL of a protein + buffer stock into every well. You will then add up to 5 μL of ligand or inhibitor.
- Different conditions will therefore end up with different final volumes. You will adjust for that volume difference during data analysis.
- Don’t add more than 5 μL DMSO to 100 μL total volume. Minimize the amount of DMSO added to the protein at each step by switching to a more concentrated stock as soon as you can, but don’t pipette less than 0.5 μL.
- The plate reader generates data in arbitrary units. This means you cannot directly compare the raw data you get from reading different plates.
Pilot 1: DNSA
- The DNSA stock will be prepared for you as follows: 1-4 mg of DNSA will be dissolved in 1 mL DMSO. In lab, we will tell you the amount used and you will need to calculate the concentration of the stock.
- Use your DNSA stock to make additional stocks in DMSO at the following concentrations: 1 mM, 100 μM and 10 μM.
- Design a pilot experiment to determine whether the protein is functional and your assay is working before you use up all of your protein setting up a full titration. Like you did in Lab 9, create a detailed table to plan your experiment. Include the following in the pilot:
- A blank
- A control for protein autofluorescence
- A control for DNSA autofluorescence
- Two experimental conditions to determine where DNSA binding is saturated: protein plus two high concentrations of DNSA. We suggest final concentrations of 20 and 40 μM DNSA.
- Measure the fluorescence of your pilot conditions on the plate reader (what are the excitation and emission wavelengths?).
- THINK ABOUT THE RESULTS FROM THE PILOT BEFORE YOU MOVE ON. If you are not confident that your assay is working, talk to your TA. You could try a higher protein concentration and/or more DNSA.
Binding of DNSA to HCAII: Full Experiment
- Design your full titration experiment such that the concentration of HCAII is constant and you add increasing concentrations of DNSA. You will then mix your decided amounts of DNSA with HCAII in a 96 well plate.
RECOMMENDATION:
In a research setting, you would run multiple pilots and perhaps full titration experiments to determine a good range of DNSA concentrations to test. We will save time by providing you a range.
We have found an optimal range for measuring DNSA binding to HCAII is approximately 50 nM – 20 μM DNSA.
- Dilute your protein in protein buffer to the desired final concentration. Pour it into a buffer trough, and use a multichannel pipette to add 100 μL protein to the appropriate wells.
- Next, pipette the varying amounts of DNSA into the designated wells – again using a multichannel.
- After pipetting components into the wells, you can mix them by gently tapping the side of the plate.
- Measure fluorescence using the plate reader. Note that no incubation time is necessary.
- Take a quick look at the data to make sure that you have reached saturation. If you don’t think you have, you may want to quickly plot [DNSA] vs. Fl. If your fluorescence is still rising dramatically at the highest [DNSA] tested, you should re-do the experiment with higher [DNSA].
Pilot 2: AZ
- Stocks of the inhibitor AZ will be provided at 1 mM. Keep in mind that you may need to dilute the stocks further. DMSO will also be provided for such dilutions.
- Design another pilot experiment to make sure that inhibition with AZ is measurable. Choose what you think would be reasonable highest and lowest [AZ] to test in this experiment. You will include these two conditions in the pilot. Create a chart like you did for the DNSA binding experiments. Include the following:
- A blank
- A control for AZ autofluorescence (you already ran controls for protein and DNSA autofluorescence so you don’t have to re-do them)
- Protein plus a saturating concentration of DNSA (this is a positive control for fluorescence signal associated with DNSA binding)
- Protein plus DNSA plus your highest [AZ]
- Protein plus DNSA plus your lowest [AZ]
- THINK ABOUT THE RESULTS FROM THE PILOT BEFORE YOU MOVE ON. If you are not confident that your assay is working, talk to your TA. You could try using less DNSA or adjust your concentration range of AZ.
Binding of AZ to HCAII: Full Experiment
- Once you determine the best concentration range, design your full AZ titration.
- This time you can create your protein stock by mixing your protein with both protein buffer and DNSA. Pipette 100 μL of the mixture per well with a multichannel pipette. Then add AZ to wells as you’ve decided.
- Measure fluorescence using the plate reader.
- Take a quick look at your data to make sure you tested reasonable concentrations of AZ.

